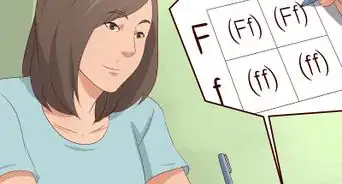X
wikiHow is a “wiki,” similar to Wikipedia, which means that many of our articles are co-written by multiple authors. To create this article, volunteer authors worked to edit and improve it over time.
This article has been viewed 12,485 times.
Learn more...
The reverse complement of a DNA sequence signifies the contents of the opposite strand in a DNA molecule. DNA molecules are constructed as such because each nucleotide has a complementary nucleotide on the other strand to which a non-covalent bond exists.
Steps
Method 1
Method 1 of 2:By Hand
Method 1
-
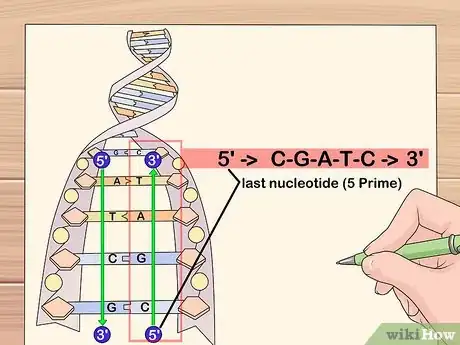
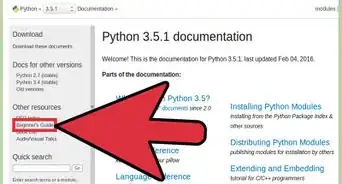
1Trace through the sequence backwards, starting from the last nucleotide in the sequence.
-
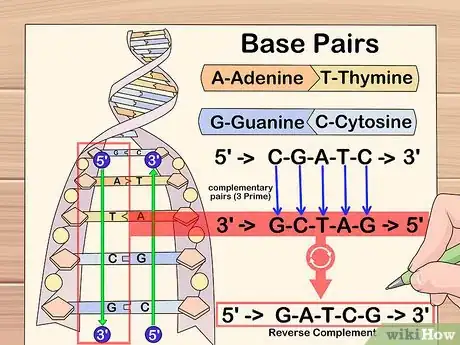
2As you pass over each nucleotide, add its complementary nucleotide to the next line, beginning the complemented string from the left-hand side of the page. Remember, guanine (G) bonds to cytosine (C) and adenine (A) bonds to thymine (T).Advertisement
Method 2
Method 2 of 2:Programmatically (Python 2)
Method 2
-
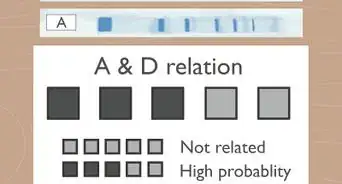
1Create or accept an input file. This article assumes that the input is in FASTA format, with a single sequence per file. The following steps also assume that all nucleotides are ATGC bases.
-
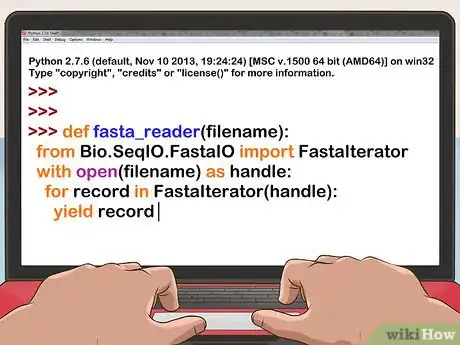
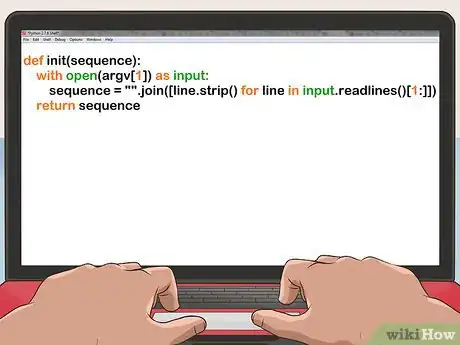
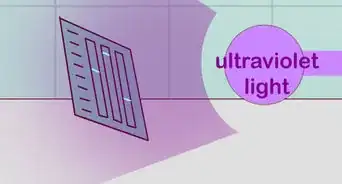
2Read in the file. For FASTA format:
- Discard the first line of the file.
- Remove all remaining newlines and other trailing whitespace.
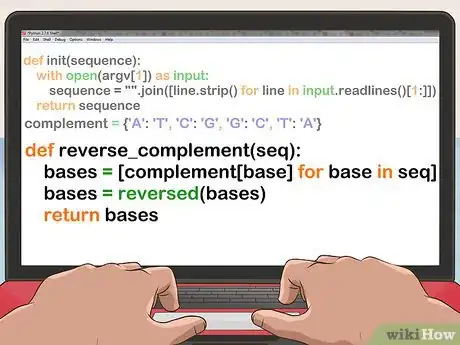
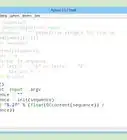
def init(sequence): with open(argv[1]) as input: sequence = "".join([line.strip() for line in input.readlines()[1:]]) return sequence
-
3Create a hash table that maps each nucleotide to its complement.
complement = {'A': 'T', 'C': 'G', 'G': 'C', 'T': 'A'}
-
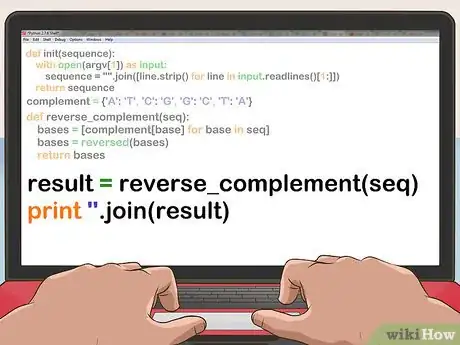
4Iterate through the sequence and use a hash table lookup to construct the complementary sequence. Reverse the resulting vector.
def reverse_complement(seq): bases = [complement[base] for base in seq] bases = reversed(bases) return bases
-
5Print the contents of the vector.=
result = reverse_complement(seq) print ''.join(result)
Advertisement
References
- StackOverflow: Reverse complement of DNA strand using Python - Source of original programmatic solution that handles non-ATGC base pairs. Solution by Gabriel. Licensed under a CC-By-SA 3.0 license.
About This Article
Advertisement