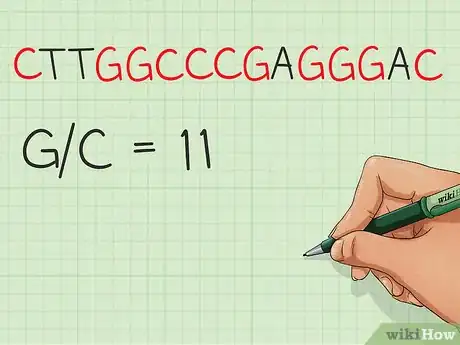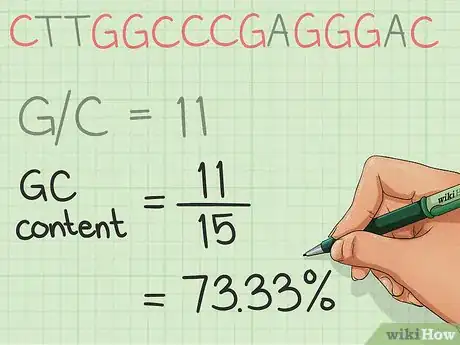X
wikiHow is a “wiki,” similar to Wikipedia, which means that many of our articles are co-written by multiple authors. To create this article, volunteer authors worked to edit and improve it over time.
This article has been viewed 9,242 times.
Learn more...
The guanine-cytosine content, or GC-content, of a DNA sequence indicates the percentage of nucleotide base pairs where guanine is bonded to cytosine. DNA with a higher GC-content will be harder to break apart.
Steps
Method 1
Method 1 of 2:By Hand
Method 1
Method 2
Method 2 of 2:Programmatically (Python 2)
Method 2
-
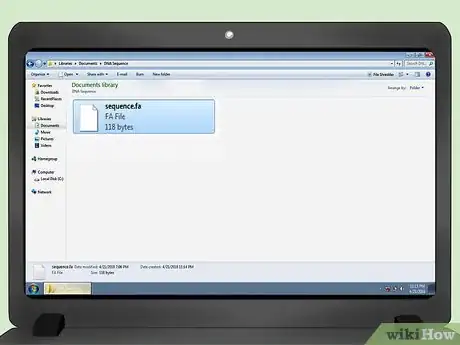
1Create or accept an input file. This article assumes that the input is in FASTA format, with a single sequence per file.
-
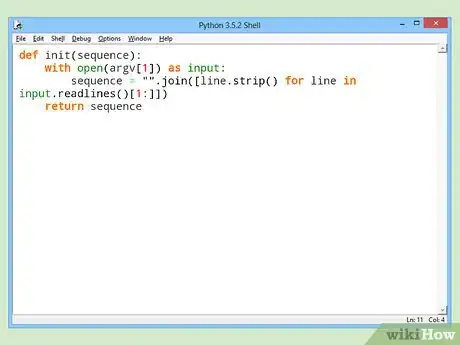
2Read in the file. For FASTA format:
- Discard the first line of the file.
- Remove all remaining newlines and other trailing whitespace.
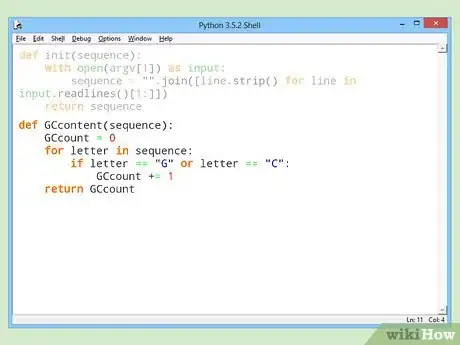
def init(sequence): with open(argv[1]) as input: sequence = "".join([line.strip() for line in input.readlines()[1:]]) return sequence
-
3Create a counter. Iterate through the data and increment your counter as you encounter any guanine or cytosine nucleotides.
-
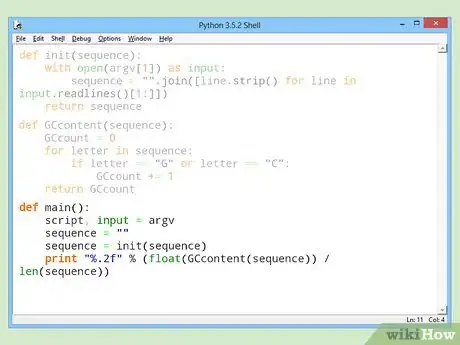
5Divide the GC count by the total length of the sequence, and output the result in percentage format.Advertisement
4
def GCcontent(sequence):
GCcount = 0
for letter in sequence:
if letter == "G" or letter == "C":
GCcount += 1
return GCcount
About This Article
Advertisement